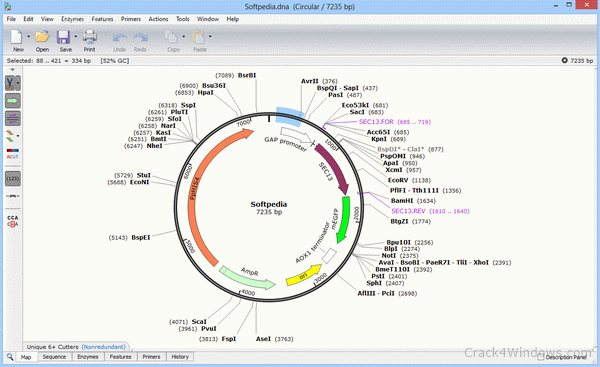
As a result, your scientists can switch entirely to SnapGene without losing data, or can continue using legacy software together with SnapGene without conflict.Īs a service to the research community, SnapGene provides tutorial videos along with a library of carefully annotated plasmids, along with guides to popular cloning methods. SnapGene supports a host of file formats.SnapGene automatically generates a record of every sequence edit and cloning procedure, so you won’t lose track of how a construct was made, even after a lab member leaves.dna files can be opened by the free cross-platform SnapGene Viewer, enabling you to share richly annotated maps and sequences with colleagues. Every DNA manipulation in SnapGene is automatically recorded, so you can see exactly what you did and retrieve the sequences of ancestral constructs.SnapGene makes your DNA manipulations easy to visualize and simulate, and alerts you to errors before they happen.These antibodies have been verified by Knockdown to. Our PER1 polyclonal antibodies are developed in Rabbit. These antibodies target PER1 in Human, Mouse and Rat samples. The software also enables documentation and sharing of data. Antibodies that detect PER1 can be used in several scientific applications, including Western Blot, Immunohistochemistry, Immunocytochemistry and ELISA. With an intuitive interface, the software enables DNA sequence visualization, sequence annotation, sequence editing, cloning, protein visualization, and simulating common cloning methods. SnapGene 6.1 is here, providing new functionality for cloning and visualizations. M, size marker (pUC19 DdeI with bands at 910, 540, 426, 409, 266, and 166 bp).SnapGene enables an easy and secure way to plan, visualize, and document everyday molecular biology procedures. (D) PCR genotyping of DNA extracted from mouse tails. (C) Southern blot analysis of DNA extracted from mouse tails. The upper band in each case represents the wild-type allele, and the lower band represents the targeted allele (see panel A). (B) Southern blot identification of the targeting event in ES cells. Alphabet: None DNA RNA Protein Nucleotide Input format: snapgene The native format used by SnapGene. Horizontal filled boxes indicate the locations of probes used for Southern blotting. (A second Kozak consensus sequence present in exon 2 could begin translation at nt 523). Filled boxes represent putative coding regions based on translation from the first ATG with a Kozak consensus sequence at nt 358 of AF050182. Open boxes represent untranslated regions. SnapGene seeks to solve these problems with well-crafted software that keeps track of the relevant variables and provides critical information at each step of a cloning simulation. Boxes represent exons, and the lines between the boxes indicate introns. Intron and exon structure is shown only for the first five exons (E1 to E5). Restriction sites introduced by the PGK-NEO cassette and used for Southern blot analysis were EcoRV (R5) and SpeI (S). The 5′ and 3′ arms of the targeting construct were ca. Make sure to use the zipped SnapGene program, other 3.2. 
Using the link instructions: Guaranteed to work on Sierra. 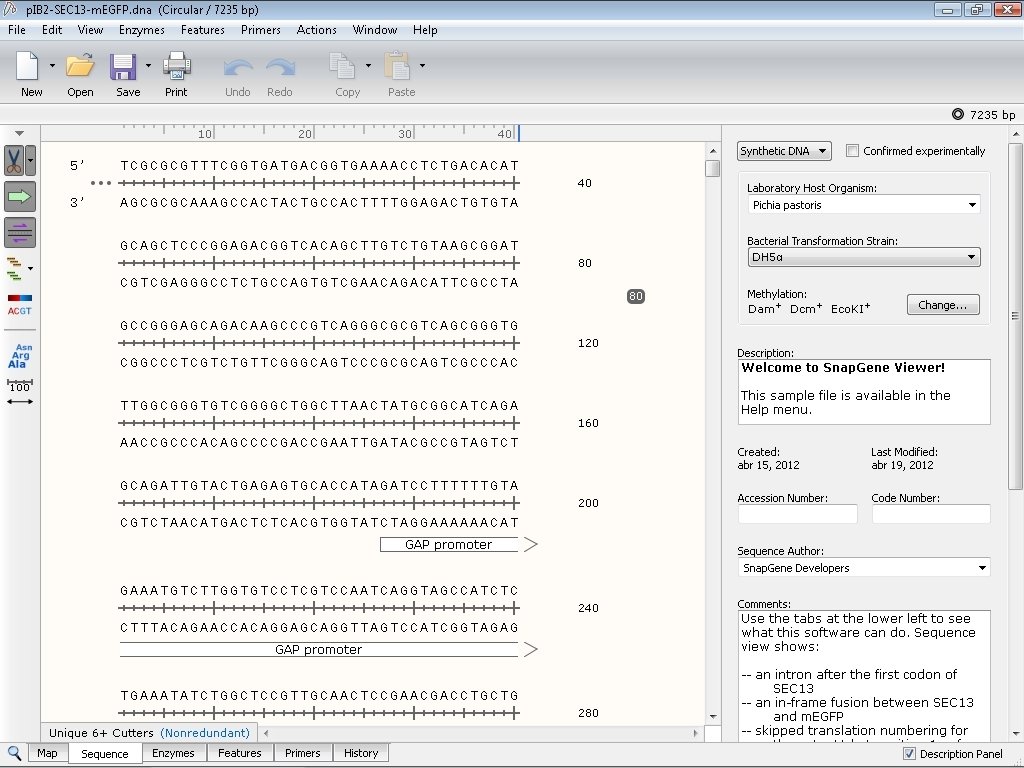
A 1.6-kb portion of the mPer3 gene was excised with EcoRI (R1), and a PGK-NEO cassette was inserted in the reverse orientation. SnapGene 3.2.1 for Mac You may need to install brew and run things from terminal, as I did. (A) Schematic representation of the mPer3 gene, the targeting construct, and the targeted allele. MPer3 targeting construct and genotyping strategies. The results demonstrate that mPer3 is not necessary for circadian rhythms in mice. Locomotor activity rhythms in mPER3-deficient mice were grossly normal, but the circadian cycle length was significantly (0.5 h) shorter than that in controls. mPer3 transcripts were rhythmically expressed in the SCN and skeletal muscle of mice homozygous for the targeted allele, but the level of expression of the mutant transcript was lower than that in wild-type controls. Rhythmic expression of mPer1 and mPer2 RNAs in skeletal muscle also did not differ between mPER3-deficient and wild-type mice. mPer1, mPer2, mCry1, and Bmal1 RNA rhythms in the SCN did not differ between mPER3-deficient and wild-type mice. Western blot analysis confirmed the absence of mPER3-immunoreactive proteins in mice homozygous for the targeted allele. To assess the role of mouse PER3 (mPER3) in the circadian timing system, we generated mice with a targeted disruption of the mPer3 gene. The basic helix-loop-helix-PAS proteins CLOCK and BMAL1 are positive regulators and drive the expression of the negative regulators CRY1 and CRY2, as well as PER1, PER2, and PER3. 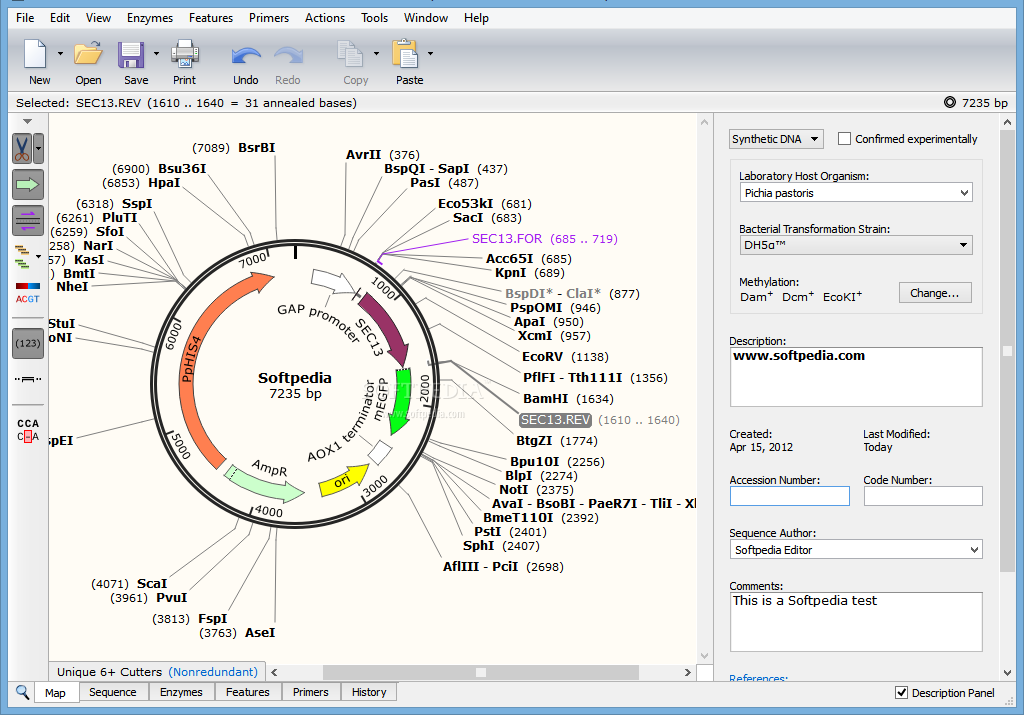
Neurons in the mammalian suprachiasmatic nucleus (SCN) contain a cell-autonomous circadian clock that is based on a transcriptional-translational feedback loop.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
